III Molecular biology techniques
III.1 Introduction
This primer outlines the chemicals and techniques used for molecular cloning, a basic technique in molecular biology. We highlight the method now called classic or traditional cloning. Since its development in 1973, it has been the most widely used cloning technique because it is easy to use, reagents are widely available, and its protocols are relatively simple. Let us define key terms before studying this method.
Recombinant DNA: A DNA molecule that contains DNA from different sources, such as two different species or two chemically synthesised DNA molecules.
Molecular cloning: The production of multiple copies of recombinant DNA by host organisms.
Transgenic or genetically modified organism: An organism that has acquired one or more foreign genes artificially.
To understand molecular cloning, we need to introduce two kinds of biomolecules: restriction enzymes and plasmids. We also need to know the techniques of gel electrophoresis and polymerase chain reaction. Gel electrophoresis separates DNA fragments based on their size and the polymerase chain reaction amplifies (makes more copies of) the DNA fragment of interest.
III.2 Restriction enzymes
Restriction enzymes belong to a class of enzymes called endonucleases that can cleave DNA. They are called endonucleases because they cleave DNA within the strand, as opposed to exonucleases that cleave the ends of DNA strands.
Restriction enzymes were discovered when scientists observed that bacteriophages (viruses that infect and replicate inside bacteria) grew in certain bacterial strains but not in others. Once the phage grew successfully inside a particular strain, it was restricted in its ability to grow on other strains. Bacteria could mount an attack on a phage that had previously grown inside a different strain.
Scientists then recognised that bacteria have enzymes that allow them to recognise foreign DNA and degrade it. They were named restriction enzymes because of their ability to restrict phage growth. The host bacterial DNA is protected from being degraded by an enzyme called methyltransferase allowing it to survive intact. Restriction enzymes and methyltransferases act together to protect a bacterium from viral invasion.
The first restriction enzymes discovered are now classified as type I, and they cleave DNA randomly. Restriction enzymes used for molecular cloning are classified as type II, and they cleave DNA at specific sites (Figure III.1). Restriction enzymes are named for the organism they are isolated from, followed by a numeral. For example, EcoRI (pronounced eco-R-one) was isolated from Escherichia coli, strain R, and it was the first such enzyme identified. BamHI (bam-H-one) was isolated from Bacillus amyloliquefaciens, strain H. Over 3600 endonucleases have now been identified, with >250 specific sites that they recognise and are able to cleave.
In many restriction enzymes, the recognition sequence is palindromic, in other words, the nucleotide sequence is the same on both strands when read in the 5’ → 3’ direction. Recognition sites can be four to eight nucleotides long. Some restriction enzymes produce blunt ends, while others produce sticky ends when cleaving DNA. Blunt ends result from symmetric cleavage on both DNA strands, whereas sticky ends result from asymmetric cleavage (Figure III.2). Fragments of DNA with sticky ends will have overhangs of unpaired bases. Sticky ends can join with other fragments that have sticky ends, if cut by the same restriction enzyme.

Figure III.1a Restriction enzyme EcoRI bound to DNA (purple and green).
David S. Goodsell, RCSB PDB, CC-BY 4.0.

Figure III.1b A cleaved DNA helix with sticky ends produced by EcoRI.
David S. Goodsell, RCSB PDB, CC-BY 4.0.

Figure III.2 Recognition sites and sticky and blunt ends produced by EcoRI and EcoRV respectively.
III.3 Plasmids
Plasmids are small DNA molecules that are extra-chromosomal, which means that they are found outside the genomic DNA of cells. Plasmids are found in bacteria, archaea and some eukaryotes such as yeast. They are circular, double-stranded, coiled or linear DNA molecules. Plasmids replicate independently of the host chromosome.
Plasmids vary from 1 to 1000 kilobase pairs, sometimes more. They can be isolated from their host cells, frozen for long periods of time, and they can move between cells. They carry genes for antibiotic resistance, toxin production, virulence, and other processes. Plasmids usually help an organism to survive challenging environments. These qualities have been found to be useful in biotechnology.
Plasmids were named by Joshua Lederberg in 1952 – a combination of ‘cytoplasm’ and ‘id’ (Latin for ‘it’) – as a general term for any extrachromosomal hereditary material in bacteria.
We will study plasmids used in bacteria, since these are the easiest and most widely used systems in biotechnology. Cloning technology has progressed to the extent that there is a large library of plasmids available for use in our experiments. It is strange to think of a library of DNA molecules, but in biotechnology terms, a library refers to a collection of plasmids stored in freezers to be thawed when needed.
Another way to store a plasmid is by keeping it in a stock of bacteria at −20°C. When the plasmid is needed, the bacteria can be thawed and cultured to increase its numbers. The plasmid can then be isolated from the bacterial population. One such plasmid library is maintained by Addgene, a non-profit plasmid repository that at last count contained > 98 000 plasmids from all over the globe.
Plasmids used in the lab contain three elements: an origin of replication, a cloning site and a selection marker (Figure III.3).
- The origin of replication (ori) is the site where replication begins. The type of ori determines whether a low number or high number of plasmid copies result when bacteria replicate.
- The selection marker is a gene that confers resistance to an antibiotic. Bacteria will be grown on an antibiotic medium. Only bacteria that contain the plasmid carrying the antibiotic-resistance gene will be able to grow on the medium. This enables scientists to select bacteria carrying the desired plasmid.
- The cloning site is where the foreign DNA is inserted into the plasmid. Plasmids are now engineered to contain a multiple cloning site (MCS). The MCS contains DNA sequences (restriction sites) that are recognised by a variety of restriction enzymes to increase its functionality.

Figure III.3 A plasmid map showing the three main elements of a plasmid.
III.4 Gel electrophoresis
Gel electrophoresis involves the application of an electric field to a mixture of biomolecules, causing them to migrate in a gel matrix. The instrument used for gel electrophoresis has a power supply for providing the electric field, an agarose gel and a UV light source.
Agarose is a polysaccharide isolated from red seaweed. Its solution is liquid at high temperature (~50°C) and solid at lower temperatures. After making an agarose solution of a specific concentration (typically between 0.7% to 2% w/v) at a higher temperature, it is cast in a tray where it cools and solidifies.
Gel electrophoresis follows this sequence of events:
- The DNA sample mixed with a dye is loaded into wells cut into the agarose gel.
- An electric field is applied for a certain amount of time.
- The dye migrates along with the DNA sample, allowing the user to follow the movement of DNA molecules.
- After a set time, the power supply is switched off and ethidium bromide dye is used to stain the DNA bands. Ethidium bromide (EtBr) has an aromatic ring structure that allows it to intercalate between DNA bases. EtBr fluoresces only when bound to DNA. It is also a known carcinogen and must be used carefully.
- The agarose gel is exposed to UV light and the ethidium bromide fluoresces, showing the DNA bands.
- Typically, a mixture of DNA strands of known molar mass, known as a ladder, is used as a standard, and the experimental DNA band size can be read from its position relative to the ladder.
DNA molecules show different mobility based on their size (molar mass), shape and charge. Other factors that affect mobility are the pore size of the gel and the amount of electric field. All these factors combine to separate the mixture into bands that migrate at a specific velocity.
Nucleic acids have a net negative charge, therefore they migrate from negative to positive electrodes with shorter fragments migrating faster than longer ones. For linear double-stranded DNA fragments, the migration distance is inversely proportional to the logarithm of the molecular mass, as shown in Figure III.4. This technique separates nucleic acids in the size range of 0.1 to 25 kilobase pairs. Watch this video for a visual guide to gel electrophoresis.

Figure III.4 An example of a readout from a gel electrophoresis experiment (left) and dependence of molecular mass (as given by number of base pairs) with migration distance (right).
III.5 Polymerase chain reaction for amplifying DNA
The polymerase chain reaction (PCR) is such an integral part of modern biology that it is difficult to remember a time prior to its existence. The idea for PCR was famously thought of in 1983 by Kary Mullis, a chemist working at Cetus Corporation, while driving to his weekend home. Mullis had been thinking of the problem of amplifying – meaning making more copies of – small quantities of DNA. He realised that small ‘primer’ complementary pieces of DNA could be used to bind to the ends of the DNA to be amplified, and the enzyme DNA polymerase could then replicate the DNA fragment. This cycle could be repeated to generate multiple copies of the DNA fragment.
After Mullis’s proof-of-principle experiment, other Cetus Corp chemists made a major improvement by choosing a DNA polymerase from the bacterium Thermus aquaticus (Taq) in the reaction. It could withstand the high temperatures of the reaction without getting degraded.
A standard PCR has the following steps.
- The DNA sequence of interest is isolated. Restriction enzymes ‘snip’ out a DNA fragment and scientists then run a gel to determine if the right fragment has been isolated. Restriction enzymes are chosen based on the sequence of the DNA fragment.
- Designing primers is the most important part of the PCR. Primers are the short (18–30 nucleotide) sequences that will bind to the DNA fragment and initiate the PCR. Primers are chemically synthesised based on the DNA fragment to be amplified, and follow certain design rules. Since the DNA fragment is a double helix, we need two primers, one for amplifying each strand. Both primers should have similar melting temperatures within a 5°C range, although this may not always be possible. There are several online programs available that can help in primer design.
- The PCR is run in a thin-walled plastic microtube with a volume of 1.5 ml. This contains the DNA fragment, primers, free nucleotides, Taq polymerase and buffer.
- The PCR involves three steps, which constitute one cycle. The reaction is carried out in a thermocycler where all these steps can be programmed (shown in Figure III.5b).
- Denaturation: The reaction temperature is raised to a point where the two strands of the DNA fragment are pulled apart or denatured. Denaturation temperatures can range from 94°C to 98°C.
- Annealing: The temperature is lowered to approximately 5°C below the primer melting temperature, allowing the primers to bind to both strands.
- Extension: The temperature is raised to 72°C. The reaction proceeds with the Taq polymerase adding free nucleotides (dNTPs) to the 3’ primer ends, extending the DNA chain. The extension time must allow enough time for the reaction to go to completion. Typical extension times are 15–30 s/kilobase. At the end of one cycle, the DNA fragment is replicated and there are four DNA molecules. Typically, PCR is run for 25 to 35 cycles to get enough amplification of the DNA fragment.
Watch this video for a demonstration of the technique.

Figure III.5a Steps in a polymerase chain reaction.
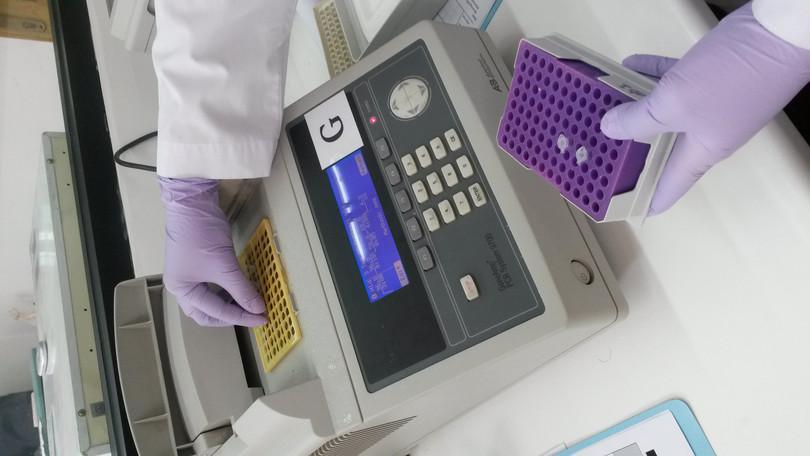

Figure III.5b A PCR machine (also called a thermocycler).
Isabella Apriyana, Wikimedia commons, CC-BY 4.0.
Conducting a molecular cloning experiment
- insert
- Also known as gene of interest, it is a fragment of DNA that is isolated from a source and inserted into a plasmid during molecular cloning.
- transformation
- The transferral of exogenous DNA into the host cell through the cell membrane, leading to the genetic alteration of the cell.
Now that we know about the molecules, gel electrophoresis and PCR, we can understand how a molecular cloning experiment is done. A typical cloning experiment has the following steps:
- Isolate the DNA/gene of interest from an organism using restriction enzymes. Two different restriction enzymes can be used for cutting the DNA strand so that it has two different ends. This DNA fragment is referred to as the insert. A gel is run to confirm that the DNA fragment of the right size has been isolated.
- Amplify this DNA fragment using the polymerase chain reaction (PCR).
- Choose a cloning vector, which is usually a bacterial plasmid. Many standard plasmids are available and can be ordered from suppliers.
- Linearise the plasmid by using a restriction enzyme to cleave its DNA. We must use the same restriction enzyme as used in step 1 so that the sticky ends in both DNA pieces are complementary.
- Combine the two DNA fragments to make recombinant DNA. The enzyme called DNA ligase joins the isolated DNA fragments to the linearised plasmid fragments.
- Introduce the recombinant plasmid into a host to clone the DNA fragment (known as transformation if the host is a bacterium).
- Make multiple copies of the DNA fragment through several rounds of host cell division on a medium containing an antibiotic. The antibiotic resistance gene on the plasmid ensures that only successfully transformed bacteria produce colonies. Untransformed bacteria will die.
Read about why cloning genes has become a standard laboratory technique.
Most of the reagents for carrying out cloning are now available as kits with defined protocols. If all instructions and reagent manufacturer recommendations are followed, there is a high probability of a successful clone being guaranteed.
The use of open-source in-silico tools for primer design, choice of restriction enzyme, plasmid and host selection allow us to optimise our protocols prior to doing any wet lab work. This ensures that all experimental steps are tested so that the chances of a successful cloning assembly are enhanced with less wastage of expensive reagents.

Figure III.6 Flow chart of molecular cloning.
III.6 Current status and the future
Molecular cloning is widely used in almost all areas of biology. The reasons for cloning a DNA molecule vary.
- Cloning allows the determination of DNA sequences of your organism of interest by replicating fragments of its DNA within a host and sequencing these fragments to assemble the genome of the organism.
- Recombinant proteins can be produced within host organisms and then purified and used for various purposes. The first such protein that was produced was insulin that was used as therapy for people with type I diabetes.
- Transgenic plants or animals can be produced. Transgenic organisms have varied purposes, for example new varieties of pest-resistant crops or animals that can produce certain proteins in their milk.
- Molecular cloning is routinely used in basic research for DNA sequence comparison and analysis and to express proteins for structure–function studies.
What has been outlined above can be called classic or traditional cloning. It is the most widely used method because it is easy to use, has simple protocols, and reagents are available. It is likely that traditional cloning will be replaced by new techniques in the coming decade. Newer protocols such as Gibson assembly and Golden Gate assembly now allow multiplexing, which means multiple fragments (or in some cases complete genes) can be cloned.
Chemical synthesis of genes is another technology that is quite advanced. Scientists have made rapid progress with oligonucleotide synthesis and are able to sequence the DNA of a variety of genes. It is now possible to synthesise longer strands of DNA/RNA molecules with shorter experimental times and reduced costs. The current technology can allow synthesis of DNA molecules between 5 and 10 kilobases. It is likely that we will soon see a combination of different technologies being used for cloning complex and large genes.
Synthetic biology has progressed rapidly in the last 15 years. One of the main drivers of synthetic biology is the International Genetically Engineered Machine (iGEM) competition. The iGEM invites teams from universities worldwide to participate in developing synthetic biology tools to solve real world problems. Many undergraduate student teams from India enter this competition every year. The iGEM foundation promotes the development of BioBricks, which are standardised inserts and plasmids that can be used by any researcher in almost a lego-like fashion. We have certainly come a long way from Cohen and Boyer’s experiment that launched this revolution.
